Block copolymers (BCPs) consist of amphiphilic molecules that can self-assemble in selective solvents and generate various types of nano- and micro-dimensional structures. The unique self-assembly process is low-cost and relatively straight forward in solution phase. The final structures have morphological diversity and complexity. These self-assembled structures have been widely used in various applications such as drug delivery, catalysis, and water purification. The self-assembly process follows a heating step for dissolution of all the components and a subsequent cooling step. Both the steps and the parameters are vital for final structural characteristics of the assembled structures. A group of scientists from two esteemed universities in Canada recently studied the effects of rate of cooling in the self-assembly process.
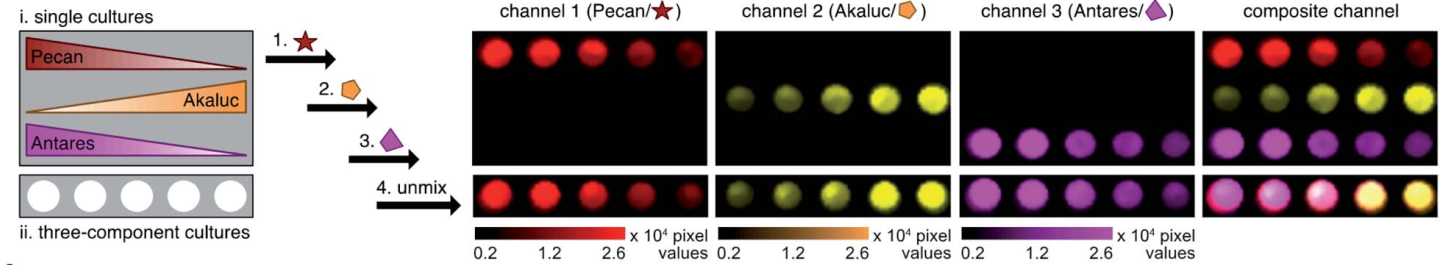
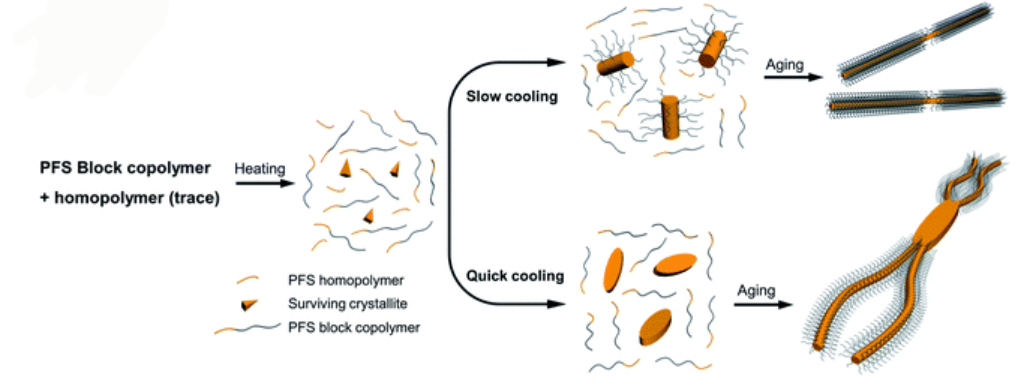
Schematic representation of how cooling rate can change the morphology of formed micellar structures using PFS BCP and homopolymers
The authors used a systematic approach to explain the influence of cooling rate on micelle morphologies for a series of PFS based BCPs. The cooling rate greatly influences the size and the shape of colloidal structures. Rapid cooling increases branching and opens a new avenue to manipulate micelle morphologies. The study finds that rapid cooling reduces crystallinity, as polymer chains do not have enough time to pack in ordered structures.
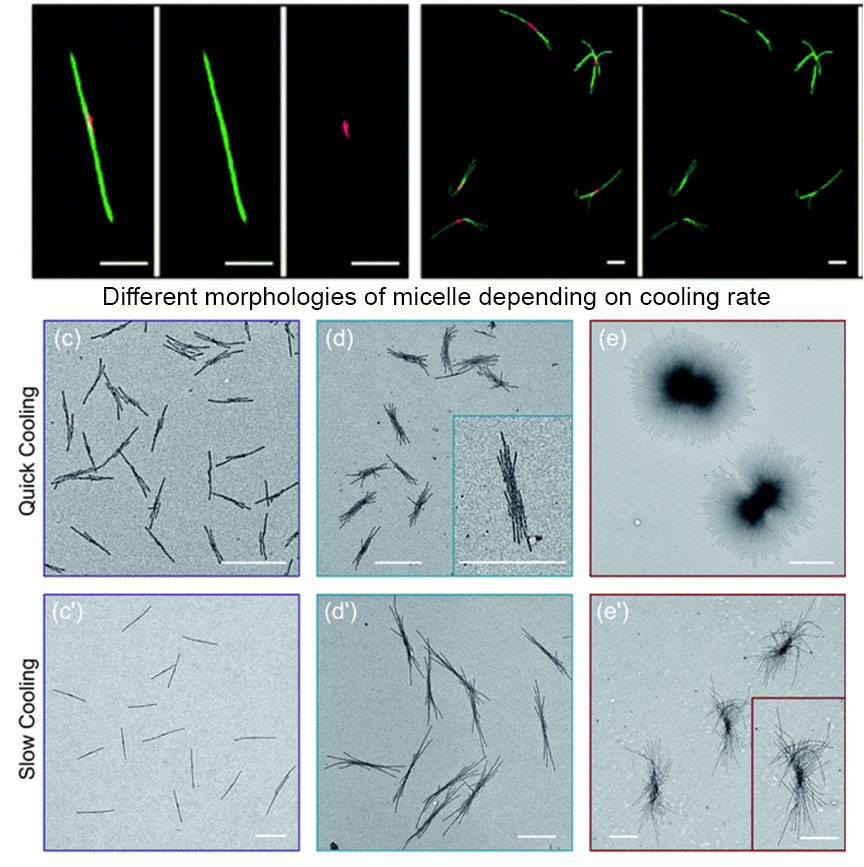
The authors standardized sample preparation protocol and then varied the cooling times, with quick cooling of 2.5 min producing flower like structures and median cooling time of 50 mins leading to the same structural features with larger size. Co-self-assembly of homopolymer BCP mixtures with variable cooling rate also shows that quick cooling generates uniform sized branched micellar structures with elongated central platelets whereas slow cooling led to a long single fiber with a dark circle platelet in the centre.
With several examples and optimization conditions, the effect of cooling in the formation of self-assembled micellar structures has been evaluated. The main outcome of this study is that the cooling rate is another parameter to manipulate crystallization-driven self-assembly and to control micelle morphologies. There exists a lot of possibilities to use the findings and apply them to generate BCPs with a crystallizable block with important optical or electronic properties.
For details, please visit the entire article at https://doi.org/10.1039/D1SC05937H
About the author:

Dr Damayanti Bagchi is a postdoctoral researcher in Irene Chen’s lab at University of California, Los Angeles, United States. She obtained her PhD in Physical Chemistry from Satyendra Nath Bose National Centre for Basic Sciences, India. Her research is focused on spectroscopic studies of nano-biomaterials. She is interested in exploring light enabled therapeutics. She enjoys food and experimenting with various cuisines, which she found resembles products/ side products of chemical reactions!