We are very pleased to introduce Siamac Fazli, Vsevolod A. Peshkov and Rustam Zhumagambetov, corresponding and first authors of the paper ‘cheML.io: an online database of ML-generated molecules‘. Their article has been very well received and handpicked by our reviewers and handling editors as one of our December HOT articles. The authors told us more about the work that went into this article and what they hope to achieve in the future. You can find out more about their article below and find more HOT articles in our online collection.
Meet the authors
 Siamac Fazli received his B.Sc. Physics degree from the University of Exeter in 2002, his M.Sc. in Medical Neuroscience from Charité University Hospital Berlin, Germany in 2004 and his Ph.D. in Computer Science from the Technical University Berlin, Germany in 2011 under the supervision of Prof. Dr. Klaus-Robert Müller. From 2011-2013 he worked as a postdoctoral researcher in the Machine Learning Group at the Technical University Berlin, Germany. In 2013, he was appointed Assistant Professor at Korea University, Seoul, Rep. of Korea. From 2016 to 2017 he worked as a Group Leader at Fraunhofer Institute for Telecommunications, Berlin, Germany. In 2018, he joined the Computer Science Department at Nazarbazev University as an Associate Professor. His current research interests include machine learning, computational chemistry and neuroscience.
Siamac Fazli received his B.Sc. Physics degree from the University of Exeter in 2002, his M.Sc. in Medical Neuroscience from Charité University Hospital Berlin, Germany in 2004 and his Ph.D. in Computer Science from the Technical University Berlin, Germany in 2011 under the supervision of Prof. Dr. Klaus-Robert Müller. From 2011-2013 he worked as a postdoctoral researcher in the Machine Learning Group at the Technical University Berlin, Germany. In 2013, he was appointed Assistant Professor at Korea University, Seoul, Rep. of Korea. From 2016 to 2017 he worked as a Group Leader at Fraunhofer Institute for Telecommunications, Berlin, Germany. In 2018, he joined the Computer Science Department at Nazarbazev University as an Associate Professor. His current research interests include machine learning, computational chemistry and neuroscience.
Dr. Vsevolod A. Peshkov received his Diploma in Chemistry in 2008 from Lomonosov Moscow State University with Prof. Nikolay V. Lukashev. In 2009, he joined the group of Prof. Erik V. Van der Eycken at the University of Leuven (KU Leuven) as a doctoral student. He defended his doctoral thesis entitled “Synthesis of nitrogen-containing medium-sized rings fused with benzene or indole through transition metal-catalyzed carbocyclizations” in 2013. He then spent one year at the University of Pittsburgh working on several medicinal chemistry projects under Prof. Peter Wipf and Prof. Donna Huryn’s direction. In September 2014, he began his independent career at Soochow University, China. In August 2018, he took on the position of Assistant Professor and Chemistry Graduate Program Director at Nazarbayev University, Kazakhstan. His research centers on a diversity-oriented synthesis (DOS) of complex heterocyclic molecules using multicomponent, one-pot and tandem strategies. In addition, his research group is active in design and synthesis of novel fluorescent organic materials and their optical properties assessment.
Rustam Zhumagambetov has received his BSc in Computer Science from the School of Science and Technology, Nazarbayev University, Kazakhstan in 2019. He is currently pursuing a Master’s degree and working as a research assistant in the Computer Science department of the School of Engineering and Digital Sciences, Nazarbayev University, Kazakhstan.
Could you briefly explain the focus of your article to the non-specialist (in one or two sentences only) and why it is of current interest?
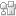
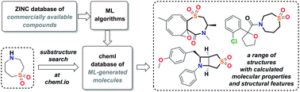
The goal of our work was to implement, validate, and compare the molecular outputs of a number of recently established machine learning algorithms for de novo molecule generation. As a result of these efforts, we created a unified database of virtual molecules in browse-able format – cheML.io. While there exists a body of literature that targets the generation of novel molecules, the audience of these works appears to be not as broad as it could be particularly because not all the researchers from the chemistry community are able to readily implement the ML algorithms described therein. That is why we decided to create our database that allows a broader audience to testify how the rapidly growing field of ML technology can be utilized for the molecular generation and in turn for the hit identification.
How big an impact could your results potentially have?
We hope that our database may provide assistance to the researchers who are interested in the chemical and biological validation of ML-generated molecules.
In your opinion, what are the key design considerations for your study?
We wanted to achieve high molecular diversity by aggregating the outcome stemming from 10 different ML frameworks into a single database. Once the database was assembled, we wanted to
couple it with a user-friendly web interface, which would allow users to browse and retrieve the data in a fast and convenient manner. Finally, we decided to provide users with the opportunity to request the generation of new molecules that could be particularly useful when a specific search leads to insufficient results.
Which part of the work towards this paper proved to be most challenging?
The most challenging part was to implement the generation on demand feature. Nevertheless, we were able to come up with the suitable solution that involves utilization of case specific training
datasets assembled through a 3-stage procedure that takes into account the structural complexity of the input motif.
What aspect of your work are you most excited about at the moment?
The generation on demand feature will allow users to contribute to the expansion of our database. We will also attempt to establish a communication channel with the users by providing them with the possibility to leave their feedback and suggestions.
What is the next step? What work is planned?
We are currently working on the establishment of new ML algorithms for molecular generation that could enhance the generation on demand feature of our database.
cheML.io: an online database of ML-generated molecules
Rustam Zhumagambetov, Daniyar Kazbek, Mansur Shakipov, Daulet Maksut, Vsevolod A. Peshkov and Siamac Fazli
RSC Adv., 2020,10, 45189-45198
DOI: 10.1039/D0RA07820D, Paper
 Submit to RSC Advances today! Check out our author guidelines for information on our article types or find out more about the advantages of publishing in a Royal Society of Chemistry journal.
Submit to RSC Advances today! Check out our author guidelines for information on our article types or find out more about the advantages of publishing in a Royal Society of Chemistry journal.
Keep up to date with our latest HOT articles, Reviews, Collections & more by following us on Twitter. You can also keep informed by signing up to our E-Alerts.