Dr. Jason Xu is an Australian Research Council (ARC) Future Fellow at School of Chemical Engineering, UNSW Sydney. He is currently leading a research group in the Cluster for Advanced Macromolecular Design (CAMD) and Australian Centre for NanoMedicine (ACN), with the focus on green and precision polymer synthesis using state-of-the-art polymerization techniques and organic chemistry tools. Dr. Xu received his BS and PhD Degrees (2007) in Polymer Chemistry from Fudan University. Following post-doctoral research in UNSW and University of Melbourne and industrial experience, he joined UNSW to develop visible light-induced living polymerization and precision polymer synthesis. He has more than 100 peer-reviewed publications in high-impact journals, attracting over 6300 citations and an h-index of 45. His areas of research interests are green chemistry and sustainable polymer synthesis, precision polymer synthesis mimicking natural perfection, advanced polymer hydrogels for strain and bio-sensors.
Dr. Jason Xu is an Australian Research Council (ARC) Future Fellow at School of Chemical Engineering, UNSW Sydney. He is currently leading a research group in the Cluster for Advanced Macromolecular Design (CAMD) and Australian Centre for NanoMedicine (ACN), with the focus on green and precision polymer synthesis using state-of-the-art polymerization techniques and organic chemistry tools. Dr. Xu received his BS and PhD Degrees (2007) in Polymer Chemistry from Fudan University. Following post-doctoral research in UNSW and University of Melbourne and industrial experience, he joined UNSW to develop visible light-induced living polymerization and precision polymer synthesis. He has more than 100 peer-reviewed publications in high-impact journals, attracting over 6300 citations and an h-index of 45. His areas of research interests are green chemistry and sustainable polymer synthesis, precision polymer synthesis mimicking natural perfection, advanced polymer hydrogels for strain and bio-sensors.
What was your inspiration in becoming a polymer chemist?
When I was still a freshman in the university, polymer chemistry was still a young and rising area at the end of 20th century in China, full of mysteries and possibilities. I was inspired by a lecture delivered by a professor in our school who is one of the pioneering researchers in polymer chemistry. He presented the amazing properties of liquid crystal polymers and foreseeable future of these materials. After that, I started to learn more about polymers and I knew polymers have already been everywhere in our life, plastics and rubbers and synthetic resins. However, there are still many things unknown for polymers, particularly for polymer chemistry. How to design and synthesize these gigantic molecules with the properties we want? This is the question, from then on, always in my mind.
What was the motivation behind your most recent Polymer Chemistry article in the Pioneering Investigators collection?
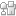
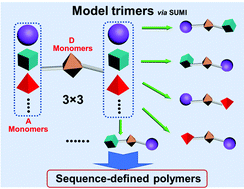
Natural biopolymers (DNA and peptides) have uniform microstructures with defined molecular weight and precise monomer sequence along the polymer chain that affords them unique biological functions. To reproduce such structurally perfect polymers through chemical approaches, researchers have proposed using synthetic polymers as an alternative. Different methodologies have been developed in the last decades. We recently proposed an emerging technology of single unit monomer insertion (SUMI), which is very similar to peptide synthesis from amino acids. SUMI can precisely prepare uniform and monodisperse alternating polymers using sequential addition of two monomers. However, the characterization of precision structure is getting harder and harder while the polymer chain increases. We therefore propose a series of short oligomers with three monomer units (trimers) to model the reaction for each step. These model trimers can provide the detailed reaction kinetics and mechanism as well as product yields, which will be the same as the reactions in long chain polymer synthesis due to the repeating monomer additions. These model trimers can also provide the reaction kinetics for copolymerization of corresponding monomers.
Which polymer scientist are you most inspired by?
There are many excellent polymer scientists I was most inspired by, such as Professors Craig Hawker and Masami Kamigaito in my mind as examples. Craig is full of very useful and bright ideas covering broad polymer area in chemistry and materials. Our recent photopolymerization technology of PET-RAFT is inspired by his pioneering work in 2012. Masami is at the forefront of polymer synthesis. His work in green polymer synthesis using renewable monomers from natural plants is fascinating.
Can you name some up and coming researchers who you think will have a big impact on the field of polymer chemistry?
This is an interesting but difficult-to-answer question. From my point of view in the specific field of polymer synthesis, there are many young and smart researchers whose research is believed to have a big impact, such as Professors Brett Fors (Cornell) and Athina Anastasaki (ETH) as examples. Their recent excellent works in photocontrolled cationic polymerization and precise polymer dispersity control are good evidence to demonstrate their potential to impact the field. Their contribution will push forward the field of polymer synthesis.
How do you spend your spare time?
I spend my spare time with my family to go out for BBQ and hiking. My daughter is currently two years old, which requires a lot of accompanying and brings so much fun to my life. Also, I like very much playing badminton and have been playing for more than 15 years. It is one of my favorite sports because it is free of any body contact different from basketball or soccer, but still requires the strength, balance and motion skills. It is therefore one of the sports anyone can keep for their whole life.
What profession would you choose if you weren’t a scientist?
I would choose my profession to be an automotive mechanic. Auto mechanic is a “precision” job like a doctor. It requires to know how all different auto parts been designed and how they work synergistically, which enables to quickly diagnose and fix the mechanical malfunction. As a mechanic, the body and mind will work all the time, which can keep the mind sharp and the body active and healthy. Actually, I hold a TAFE Auto mechanic certificate and always had a plan to run a workshop. What I need now is the financial support from some potential investors (kidding!).
Read Jason’s full article now for FREE until 17 November!
Also check our the work of our other Pioneering Investigators here
Sequence-defined polymers have garnered increasing attention in a broad range of applications from materials engineering to medical science. Reversible addition–fragmentation chain transfer single unit monomer insertion (RAFT SUMI) technology has recently emerged as a powerful tool for sequence-defined polymer synthesis, which utilizes sequential monomer radical additions occurring one unit at a time to assemble olefins into uniform polymers. The strategy of employing alternating additions of electron-donor and acceptor (D–A) monomers can be used to prepare long chain sequence-defined polymers by the RAFT SUMI technique. However, considering both terminal and penultimate unit effects, complex radical reaction kinetics can result from various monomer addition orders particularly if three or more different families of vinyl monomers are used to build diverse sequences. Simplifying reaction processes and establishing reaction kinetics will be critical for effective synthesis of sequence-defined polymers. Herein, a series of model trimers containing D–A–D and A–D–A triads was thus produced from four families of α,β-disubstituted vinyl monomers (N-phenylmaleimide, fumaronitrile and dimethyl fumarate and indene). Such trimers presented distinct synthesis kinetics (reaction rate and yield). These model trimers and their kinetics data are able to provide full guidance for the synthesis of long chain discrete polymers using sequential and alternating RAFT SUMI processes.
About the Webwriter
 Simon Harrisson is a Chargé de Recherche at the Centre National de la Recherche Scientifique (CNRS), based at the Laboratoire de la Chimie des Polymères Organiques (LCPO) in Bordeaux, France. His research seeks to apply a fundamental understanding of polymerization kinetics and mechanisms to the development of new materials. He is an Advisory Board member for Polymer Chemistry. Follow him on Twitter @polyharrisson
Simon Harrisson is a Chargé de Recherche at the Centre National de la Recherche Scientifique (CNRS), based at the Laboratoire de la Chimie des Polymères Organiques (LCPO) in Bordeaux, France. His research seeks to apply a fundamental understanding of polymerization kinetics and mechanisms to the development of new materials. He is an Advisory Board member for Polymer Chemistry. Follow him on Twitter @polyharrisson