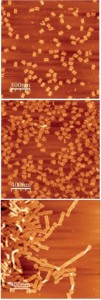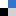
Schematic of DNA translocation in a glass capillary. Image taken from Soft Matter, 2012, doi:10.1039/C2SM25346A
Detecting single strands of DNA is a tricky business. One way in which it can be done is by creating a pore and sensing the DNA strand as it passes through the pore. This is what Ullrich Keyser from the University of Cambridge, UK and his group have been doing.
Keyser forms nanopores with diameters of less than 100nm from glass capillary tubes (the diameters can be as small as 20nm). These nanocapillaries act as single molecule sensors. Using electrophoresis, the negatively charged DNA is pulled towards a positively charged electrode inside the capillary, in a process known as DNA translocation. As the strand enters the capillary the resistance across the capillary pore changes, allowing the DNA to be detected. The change in ionic current is dependent not only on the presence of a DNA strand, but also on its folded state. This method offers a simple and cost-effective method for the detection of single molecules and for DNA sequencing.
Full control over DNA translocation can be achieved using optical trapping, where the DNA is attached to a colloidal particle, held in place by an optical trap. The DNA can then be moved in and out of the capillary pore at will. Using this method, Keyser and his group have measured the capture force due to the electric field acting on the DNA. Their results show that the DNA capture force is linearly dependent on the number of strands captured in the capillary.
Whether using glass capillaries, or pores formed in silicon nitride membranes via focussed ion beam milling, control over the exact shape and functionality of the nanocavity can be problematic. Keyser has taken DNA detection yet another step further by using the DNA itself to create the nanopore. The shape into which a DNA strand folds can be controlled in a a process known as DNA origami; the DNA is synthesised such that it will self-assemble into a pre-designed three-dimensional shape. Using this origami, it is possible to design and fabricate virtually any nanosized shape that you want.
Keyser designed the DNA so that it folded into a funnel like shape with a long tail. This structure was then pulled through a pore in a silicon nitride membrane to form a hybrid nanopore with a diameter of 7.5nm. The assembly of the hybrid pore is robust and easily reversible. These DNA/silicon nitride pores have been successfully used to detect single strands of DNA. The hybrid nanopores offer a novel way to change the size, shape and functionality of pores.
Relevant papers in SoftMatter:
Chen, Q. et al., How does a supercoiled DNA chain pass through a small conical glass pore? Soft Matter, 2012, Advanced Article.
Geerts, N., Eiser, E., DNA functionalized colloids: Physical properties and applications. Soft Matter, 2010, 6, 4647-4660.
Kim, K. N., et al., Comparison of methods for orienting and aligning DNA origami. Soft Matter, 2011, 7, 4636-4643.