Early career researchers in chemical biology are not only shaping the future of their field, they are also redefining what scientific leadership looks like. This September, early career researchers (ECRs) gathered at Imperial College London for the second Chemistry Biology Interface Community (CBIC) Early Career Researcher Leadership Retreat, a three-day residential event designed to strengthen leadership skills, encourage collaboration, and build a supportive professional network.
Organised by CBIC Council members Anna Barnard, Gavin Miller and Andrew Truman, with local coordination support from Imperial’s Institute of Chemical Biology, the retreat brought together a diverse group of participants from across the UK and beyond. The event was sponsored by RSC Chemical Biology, and supported with a grant from the RSC’s Chemistry Biology Interface Community, with RSC staff joining throughout the programme to share resources and highlight opportunities available for ECRs across publishing, funding, and professional development.

Day 1: Research, publishing, and RSC support for early career researchers
The retreat opened with a warm welcome from the organisers and an introduction to the programme’s aims: to provide a dedicated space where early career researchers could reflect on their professional journeys and exchange ideas on leading research teams effectively.
The first morning included an invited talk from Professor Ed Tate (Imperial College London), who shared insights into advances in chemical biology and the importance of interdisciplinary collaboration. His talk set a motivating tone for the retreat.
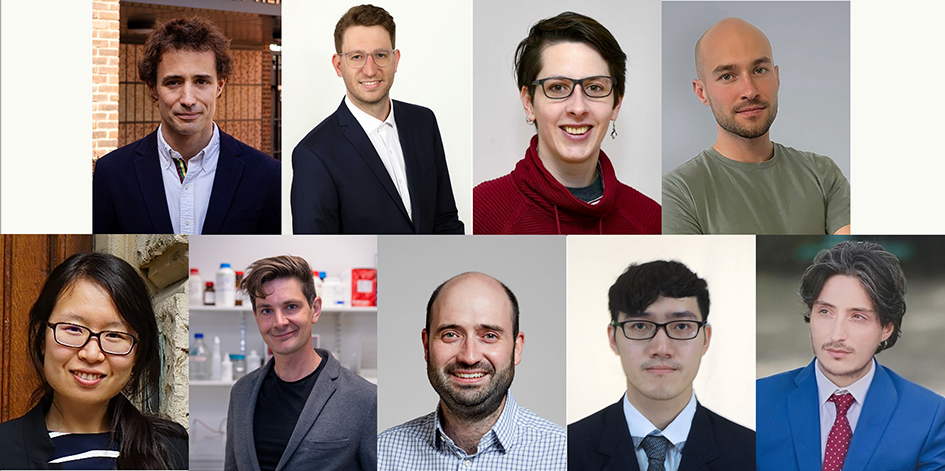
A highlight of the opening day was a session of flash presentations, where each participant delivered a short overview of their research area. These rapid-fire talks helped participants learn about one another’s scientific interests, sparking discussions that continued throughout the event.
Later, the session focused on research and publishing, with the RSC Chemical Biology Executive Editor, Anna Rulka, presenting an overview of publishing with the Royal Society of Chemistry and the support available to early career authors. Delegates heard about opportunities to contribute to themed issues, editorials, and community-led features, as well as guidance on navigating the publication process.
These discussions sparked valuable conversations around scientific communication, authorship, and how ECRs can raise the visibility of their work.
Throughout the day, participants exchanged insights on research leadership, group management, and the evolving landscape of chemical biology research, setting the tone for an open, collaborative retreat.

Dr Anna Rulka’s presentation
Day 2: Leadership skills and industry insights
The second day centred on leadership and industry engagement, with sessions designed to help participants reflect on how they lead, communicate, and collaborate.
Elizabeth Adams from Scafell Coaching facilitated an interactive morning workshop exploring what effective leadership looks like in research environments. Drawing on her experience, she encouraged participants to consider the importance of positive team culture, clear communication, and wellbeing in academic settings.
Through group exercises, attendees developed team charters, discussed approaches to conflict management, and shared practical strategies for workload balance and mentoring students. The conversations highlighted that leadership in science goes beyond technical expertise, it’s also about fostering inclusion, trust, and collaboration.
Collaboration was a recurring theme throughout the day. Delegates explored how to find suitable research partners, set clear expectations, and ensure fair recognition using different frameworks. Reviewing each other’s online profiles also helped participants reflect on how to represent their work effectively to potential collaborators and employers.
In the afternoon, participants heard from Dr Nadia Luheshi of AstraZeneca, whose talk offered an inspiring view of career pathways in industry. With experience spanning immunology, mRNA research, and oncology, she spoke about her transition from academia to industry and shared insights into open innovation, target validation, and the future of cancer research.
Nadia’s talk also highlighted how AI, data science, and digital health are transforming research and diagnostics, reinforcing how chemical biologists can contribute to advancing sustainable, patient-centred innovation.

Dr Nadia Luheshi
Day 3: Research sharing and reflection
The final day of the retreat returned to a focus on research, with each participant delivering a “chalk talk”: a concise five-minute presentation showcasing new data, challenges, or ideas from their lab, followed by a short discussion.
These sessions were energetic and collaborative, reflecting the diversity of expertise represented in the room. Participants provided constructive feedback, discovered shared interests, and even identified potential collaborations to pursue after the retreat.
The CBIC Programme Manager joined the final session to share upcoming opportunities for ECRs, including an RSC Funding Workshop on establishing independent research careers. As the retreat drew to a close, many participants reflected on how the experience had strengthened their confidence, expanded their professional networks, and provided new tools for leadership.

Prof. Ed Tate at his plenary
Strengthening the early career chemical biology community
The CBIC Early Career Researcher Leadership Retreat aims to become a cornerstone event for the chemical biology community. Building on the success of the inaugural 2023 meeting in Newcastle, this second edition continued to provide a supportive and intellectually stimulating environment for emerging leaders in the field. The quality of the ECRs at the event indicates that the future is bright for chemical biology research in the UK.
By combining scientific discussion with leadership training and peer-to-peer learning, the retreat underscored the importance of nurturing the people who will shape the next generation of chemical biology research. Feedback from delegates highlighted the value of the programme’s mix of sessions, which balanced professional development with time for reflection and collaboration.
For the RSC, events like this are an important way to connect with early career researchers, listen to their experiences, and ensure that Royal Society of Chemistry continues to meet their evolving needs—from publishing support and career development resources to funding opportunities and community networks.
The RSC extends sincere thanks to the organisers, speakers, and participants for delivering another inspiring retreat. Their enthusiasm, openness, and collaborative spirit reflect the best of the chemical biology community.

Read more: RSC CBIC Leadership Retreat Empowers Next-Gen Chemical Biologists | Imperial News | Imperial College London
Comments Off on Building connections and confidence: the 2nd CBIC Early Career Researcher Leadership Retreat