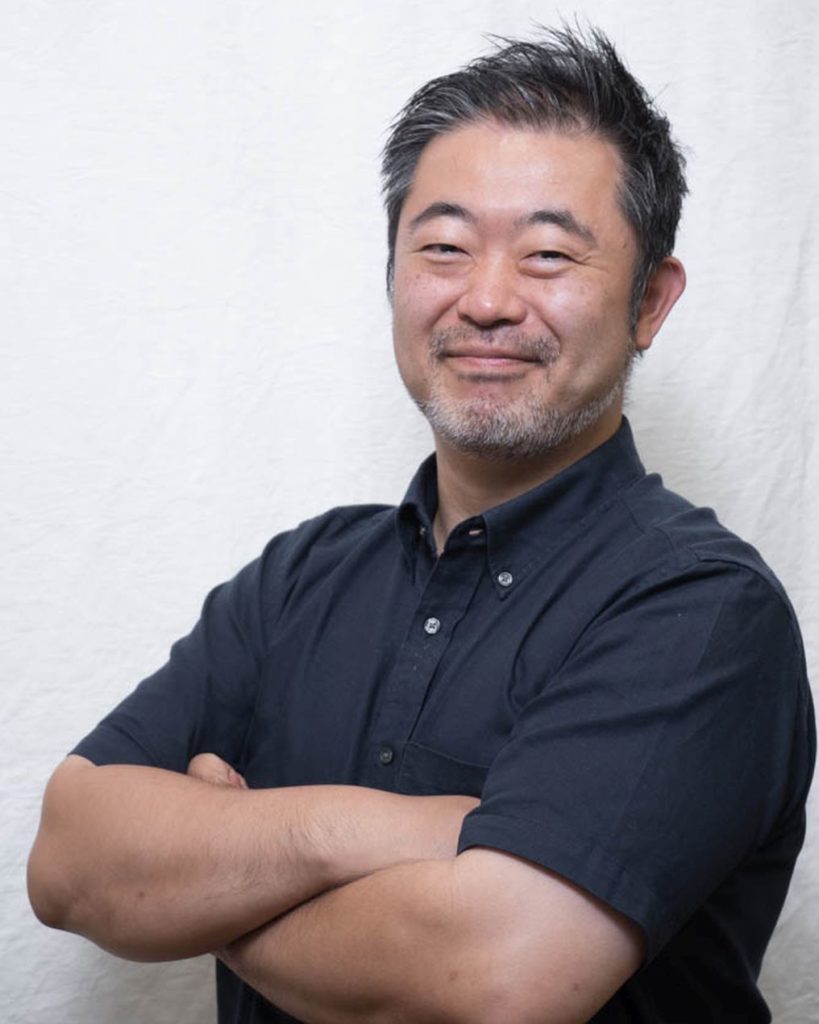
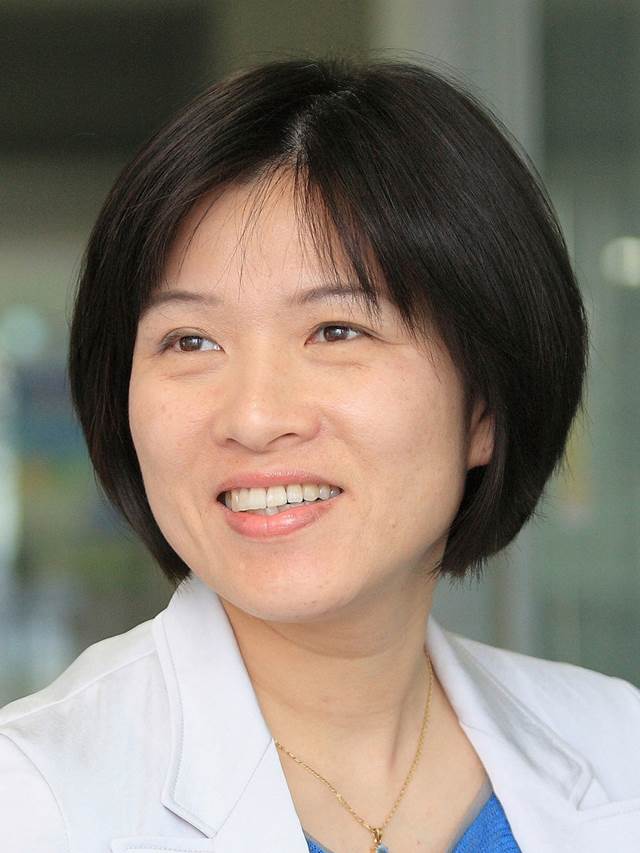
We are delighted to welcome Professors Keisuke Goda, Sindy Tang & Yi-Chin Toh to the Lab on a Chip Advisory Board.
Professor, Department of Chemistry
University of Tokyo, Japan
Keisuke Goda is a professor in the Department of Chemistry at the University of Tokyo, an adjunct professor in the Institute of Technological Sciences at Wuhan University, and an adjunct professor in the Department of Bioengineering at UCLA. He obtained his B.A. from UC Berkeley summa cum laude in 2001 and his Ph.D. from MIT in 2007, both in physics. At MIT, he worked on the development of gravitational-wave detectors in the LIGO group which led to the 2017 Nobel Prize in Physics. After several years of work on high-speed imaging and microfluidics at Caltech and UCLA, he joined the University of Tokyo as a professor. His research group focuses on the development of serendipity-enabling technologies based on molecular imaging and spectroscopy together with microfluidics and computational analytics to push the frontier of science. He currently leads Serendipity Lab, a global network of scientists who aim to realize Louis Pasteur’s statement “Chance favours the prepared mind”. He has published >300 papers, filed >30 patents, and received numerous awards and honours such as Japan Academy Medal and JSPS Prize. He is a fellow of RSC and SPIE.
Keisuke was the recipient of the Lab on a Chip & Dolomite Pioneers of Miniaturization Lectureship Award 2021, and is a Thought Leader for our on-going AI in Microfluidics thematic collection.
Find out more on the GODA LAB website and follow @ut_godalab on Twitter
Here are a selection of Keisuke’s Lab on a Chip papers below:
- Deep imaging flow cytometry
- Are droplets really suitable for single-cell analysis? A case study on yeast in droplets
- AI on a chip
Sindy Tang
Associate Professor of Mechanical Engineering and by courtesy of Radiology (Precision Health and Integrated Diagnostics)
Stanford University, USA
Sindy KY Tang is the Kenneth and Barbara Oshman Faculty Scholar and Associate Professor of Mechanical Engineering and by courtesy of Radiology (Precision Health and Integrated Diagnostics) at Stanford University. She received her Ph.D. from Harvard University in Engineering Sciences under the supervision of Prof. George Whitesides. Her lab at Stanford works on the fundamental understanding of fluid mechanics and mass transport in micro-nano systems, and the application of this knowledge towards problems in biology, rapid diagnostics for health and environmental sustainability. The current areas of focus include the flow physics of confined micro-droplets using experimental and machine learning methods, interfacial mass transport and self-assembly, and ultrahigh throughput opto-microfluidic systems for disease diagnostics, water and energy sustainability, and single-cell wound healing studies. She was a Stanford Biodesign Faculty Fellow in 2018. Dr. Tang’s work has been recognized by multiple awards including the NSF CAREER Award, 3M Nontenured Faculty Award, the ACS Petroleum Fund New Investigator Award, and invited lecture at the Nobel Symposium on Microfluidics in Sweden.
Tang is the Kenneth and Barbara Oshman Faculty Scholar and Associate Professor of Mechanical Engineering and by courtesy of Radiology (Precision Health and Integrated Diagnostics) at Stanford University. She received her Ph.D. from Harvard University in Engineering Sciences under the supervision of Prof. George Whitesides. Her lab at Stanford works on the fundamental understanding of fluid mechanics and mass transport in micro-nano systems, and the application of this knowledge towards problems in biology, rapid diagnostics for health and environmental sustainability. The current areas of focus include the flow physics of confined micro-droplets using experimental and machine learning methods, interfacial mass transport and self-assembly, and ultrahigh throughput opto-microfluidic systems for disease diagnostics, water and energy sustainability, and single-cell wound healing studies. She was a Stanford Biodesign Faculty Fellow in 2018. Dr. Tang’s work has been recognized by multiple awards including the NSF CAREER Award, 3M Nontenured Faculty Award, the ACS Petroleum Fund New Investigator Award, and invited lecture at the Nobel Symposium on Microfluidics in Sweden.
Sindy has been a committed member of the Lab on a Chip 2022 Commissioning Panel
Find out more on the Tang Lab website and follow @TangSindy on Twitter
Here are a selection of Sindy’s Lab on a Chip papers below:
- Exponential magnetophoretic gradient for the direct isolation of basophils from whole blood in a microfluidic system
- Confinement and viscosity ratio effect on droplet break-up in a concentrated emulsion flowing through a narrow constriction
- Optofluidic ultrahigh-throughput detection of fluorescent drops
Yi-Chin Toh
Associate Professor, School of Mechanical, Medical and Process Engineering,
Queensland University of Technology, Australia
Yi-Chin Toh is a Future Fellow and Associate Professor at the Queensland University of Technology. She obtained her B.Eng in Chemical Engineering and Ph.D in Bioengineering from the National University of Singapore in 2001 and 2008 respectively. She did her post-doctoral training at the Massachusetts Institute of Technology in 2008 under Professor Joel Voldman’s guidance. Before joining QUT, she led an independent research group as an Assistant Professor at the Department of Biomedical Engineering, National University of Singapore.
Her research interest is in engineering multi-scale tissue models to mimic complex biological interactions during human development and diseases, as well as translating them into scalable platforms for disease modeling and drug testing applications. Dr Toh is a recipient of the Australia Research Council Future Fellowship, National University of Singapore Research Scholarship, A*STAR Graduate Scholarship and A*STAR International Fellowship.
Yi-Chin has been a committed member of the Lab on a Chip 2022 Commissioning Panel
Find out more on the MicroTE Lab website and follow @MicroTElab on Twitter
Here are a selection of Yi-Chin’s Lab on a Chip papers below:
- Integration of a microfluidic multicellular coculture array with machine learning analysis to predict adverse cutaneous drug reactions
- Self-aligning Tetris-Like (TILE) modular microfluidic platform for mimicking multi-organ interactions
- A liver-immune coculture array for predicting systemic drug-induced skin sensitization
Please join us in welcoming our new Advisory Board members and we look forward to continuing to work with them on Lab on a Chip!