 Advances in the development of genomic engineering have unprecedented potential to make significant impacts on the future of health and research. Since their development, genetic editing tools have become a major hot topic not only within the scientific community but also with the general public. Precise and facile methods of modifying gene sequences provide us with the ability to understand and even cure some of our deadliest genetic diseases however, this has also raised ethical questions as to what can (and perhaps should) be achieved.
Advances in the development of genomic engineering have unprecedented potential to make significant impacts on the future of health and research. Since their development, genetic editing tools have become a major hot topic not only within the scientific community but also with the general public. Precise and facile methods of modifying gene sequences provide us with the ability to understand and even cure some of our deadliest genetic diseases however, this has also raised ethical questions as to what can (and perhaps should) be achieved.
In their most basic form, genome editing technologies are proteins that can cleave DNA in a site specific manner and cause alterations such as inactivation of specific genes, correction of mutated sequences or insertion of intact genes into a target DNA sequence. This cumulates into a downstream phenotypic change within the cell that not only allows biologists to study normal gene function but also carries tremendous therapeutic relevance.
A recent Tutorial Review by Debojyoti Chakraborty and Souvik Maiti published in Chemical Society Reviews describes the development, evolution and future of genetic editing technologies and molecular tools such as zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), site specific recombinases, and clustered regularly interspaced short palindromic repeats (CRISPR). Of those listed, the CRISPR-Cas9 system is by far the most precise, efficient, inexpensive and easy-to-use tool for genetic manipulation and has been proven to work on virtually any living cell in almost any organism. In the short amount of time since its development, CRISPR-Cas9 has reinvented the way in which scientists are carrying out genetic research and approaching gene therapy.
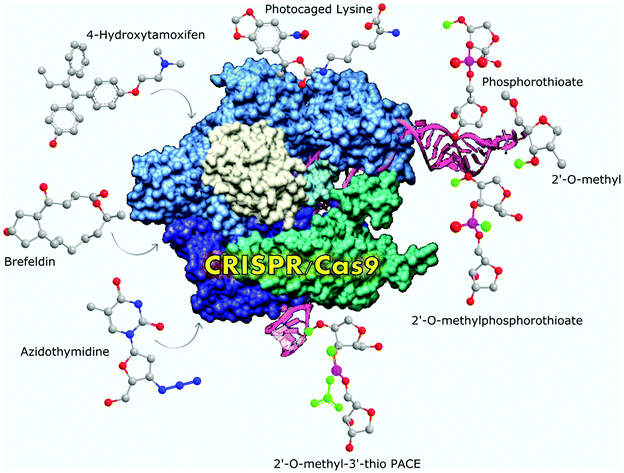
CRISPR-Cas9 is distinct from conventional gene therapies which are often only capable of crudely placing genetic material at a random location within the cell. The idea behind CRISPR-Cas9’s mechanism and its precision is derived from innate bacterial immune systems which use arrays of short repeats in the microbial genome, termed CRISPR, to protect themselves from foreign genetic material. The CRISPR-Cas9 system is made up of the Cas9 protein that snips the DNA strands and an RNA component that directs the complex to the correct sequence. The cell’s natural machinery then repairs the cut but it is error-prone and will likely make a mistake that will result in termination of that gene’s function. This allows scientists to determine the gene’s role in cellular function. An interesting example of this was recently published in Chemical Science by Peter Leadlay of Cambridge University and Yuhui Sun of Wuhan University School of Pharmaceutical Sciences and describes the application of this genomics-based approach in determining the biosynthetic pathway of thiotetronate antibiotics. Alternatively, the cut can be repaired using a DNA template provided by the researchers to edit nearly any sequence at any site in the genome.
The CRISPR-Cas9 system has been demonstrated in numerous studies to be a highly efficient tool for gene-editing and has resulted in significant advancements within the field of genomics. There are however, as with any new technology, limitations and gaps in our understanding that need to be addressed before clinical applications can be considered. Unsurprisingly, a lot of research is being focused on tackling these issues. The potential to rewrite the human genome to alter disease states, perhaps enhance our ability to fight infectious diseases and even alter pathways involved in aging is incredible but will inevitably come with a spectrum of ethical, social and intellectual challenges. In any case, these are exciting times and these inventions stand to change the future of medicine.
To find out more see:
CRISPR/Cas9: a historical and chemical biology perspective of targeted genome engineering
Amrita Singh, Debojyoti Chakraborty and Souvik Maiti
DOI: 10.1039/C6CS00197A
A genomics-led approach to deciphering the mechanism of thiotetronate antibiotic biosynthesis
W. Tao, M. E. Yurkovich, S. Wen, K. E. Lebe, M. Samborskyy, Y. Liu, A. Yang, Y. Liu, Y. Ju, Z. Deng, M. Tosin, Y. Sun and P. F. Leadley
DOI: 10.1039/C5SC03059E