A peek at who we are, how we run the site, and how we manage data quality.
What is ChemSpider and who runs the service?
ChemSpider is one of the largest chemical databases in the world, containing data on over 65 million chemical structures. This data is freely available to the public at ChemSpider.com, a website published by the Royal Society of Chemistry.
How does the Royal Society of Chemistry support ChemSpider?
ChemSpider.com is an independent service that does not rely on direct or research grant funding. The Royal Society of Chemistry supports the website using the surplus generated by our publishing activities, allowing us to provide a sustainable and reliable service. We also generate revenue from advertising and by providing paid for web services, such as our APIs, for non-academic users. These activities help keep ChemSpider financially sustainable and help support our server costs, staff hours and development.
These services enable us to make the site available free anyone in the world, and we reached over six million unique users in 2017. These users range from school students looking for help with their homework, to researchers working in academia and industry, to general users who want to keep their chemical knowledge up to date. They come from every continent except Antarctica, and just about every country on Earth.
What goes into ChemSpider?
ChemSpider data comes from the chemical sciences community itself – submitted by researchers, databases, publishers, chemical vendors and many more.
We have two main inclusion criteria for ChemSpider data:
- Machine readability – Depositors must provide structures in a machine-readable format, typically a .mol file that is interpretable by InChI – the open-source chemical structure representation algorithm.The .mol format describes how a compound is arranged, atom-by-atom and bond-by-bond. This means that it can only accurately depict small molecules with defined structures. For ChemSpider, “small” means structures up to 4000 daltons, including short peptides, oligonucleotides, and other structures. Large proteins, extended crystal lattices or long nucleotides are too big to describe sensibly in ChemSpider, but are available from other databases suited for larger molecules.
We also only accept ‘defined structures’ – compounds with exact chain lengths, fully expressed functional groups, and integer bond orders – due to the requirement to describe every heavy atom in a molecule. This means we can only accept structures for which we can generate a valid InChI.
Most ChemSpider structures are organic molecules. However, we do accept some inorganic and organometallic compounds, with specific methods for curating these.
- Real compounds – We do not accept virtual or prophetic compounds.
As far as possible, we only accept compounds that have been synthesised or isolated in physical form. This means we do not accept transition states, theoretically predicted compounds, virtual compounds from vendors or prophetic compounds from patents.
Who are our data sources?
We have received data from almost 250 unique data sources, including data from chemical vendors, specialist databases, individuals, research groups and publishers. These sources cross the breadth of the chemical sciences – including biochemistry, pharmacology and toxicology, natural products, spectroscopy and crystallography. Each ChemSpider record includes links to all of the data sources for the compound, enabling users to find and to check the provenance of the data.
Our data source list is continually changing, as we find new sources of data to add and remove outdated or low-quality data sources.
We no longer accept data from other data aggregators. We have taken this step to match our quality requirements with other databases and reduce the propagation of algorithmically generated errors that can arise from prophetic sources. One example of this is Chessboardane, which originated from an optical structure recognition program interpreting a data table contained within a patent as a chemical structure. The result was an 81-carbon grid structure, erroneously identified as a complex cyclic alkane, which was deposited in a public repository and shared between multiple aggregators.
Because of this, we only seek data directly from the original sources, where we have greater certainty about the data’s provenance and accuracy, and are working to curate legacy data still within ChemSpider.
Because of examples like Chessboardane, we are cautious about accepting data from text-and-data-mined sources that depositors have programmatically extracted from text or encoded images in patents or scientific literature. After review, we have added some of the highest quality data mined sources. We will continue to review potential new data-mined sources on a case-by-case basis to ensure that their data meet our quality standards.
Automated filters
A manual check of every one the 65 million records in ChemSpider would take an individual more than 600 years to complete working round the clock – even if we only invested five minutes of curation time per record.
Instead, we run each deposition through a series of automated filters to pick out unsuitable structures, such as those with incorrect valences, unbalanced charges, or missing stereochemistry. In addition to structure filters, we also apply basic name and synonym filtering and regularly review the processed files so that we can improve our filters.
We have provided a simplified overview of this process below, and will provide a more detailed description of our filters in a separate blog post:
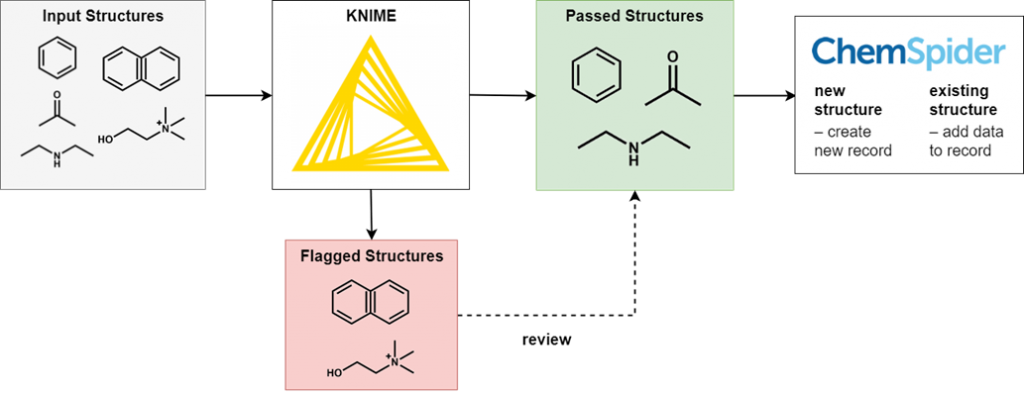
Curation by ChemSpider staff
ChemSpider is run by a small team of full-time curators, who work to add new compounds, remove errors, and respond to user feedback. Our staff have extensive experience of both chemical data and practical chemistry, with backgrounds in fields such as organic synthesis and art conservation, and a wealth of experience working on other Royal Society of Chemistry databases, such as The Merck Index* Online and Analytical Abstracts.
Community curation
Because we cannot review every record ourselves, we really appreciate comments or corrections from our users. The easiest way to help us improve ChemSpider is to leave feedback or email us when you spot an error. We try to act on user feedback within a few days – sooner for simpler queries. Please let us know if you find an error by leaving a comment on the relevant ChemSpider record, or by emailing us (chemspider@rsc.org).
Users wishing to get more involved can directly deposit structures and curate synonyms related to their research or work, without having to email the ChemSpider team.
We are extremely grateful for all the contributions our community curators have made over the years.
Keep using and contributing to ChemSpider
To access information on over 65 million chemical structures, go to ChemSpider.com, which is fully searchable by structure, name, or advanced query, from any device, anywhere, for free.
To deposit data, tell us about an error, become a curator, or for any other query, please do not hesitate to email us at chemspider@rsc.org
*The name THE MERCK INDEX is owned by Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc., Whitehouse Station, N.J., U.S.A., and is licensed to The Royal Society of Chemistry for use in the U.S.A. and Canada.

