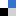
Differentiating quadruplexes from duplexes using silver nanoparticles and SERS
A promising drug target and ligand-binding site, G-quadruplexes form via Hoogsteen hydrogen bonding in guanine bases. These quartets are rich in guanine and are stabilized through π-π stacking interactions and a cation such as potassium or sodium. As researchers explore new compounds to target this region to treat diseases such as cancer, the structure of these quadruplexes and their response to ligands compared to standard DNA remains understudied.
Researchers at the University of Strathclyde have used the sensitivity of surface-enhanced Raman spectroscopy (SERS) to detect the formation of G-quadruplexes in the presence of stabilizing ligands and silver nanoparticles. They successfully differentiated the quadruplex DNA structure from the duplex based on the SERS signal, and demonstrated the capability of SERS in identifying higher order DNA structures. To read more about the qualitative detection of these structures, click the link below. It will be free to read until June 30.
K. Gracie, V. Dhamodharan, P. I. Pradeepkumar, K. Faulds and D. Graham
Analyst, 2014, Advance Article